Key Points
-
Whole-genome single-nucleotide polymorphism (SNP) mapping to determine pharmacogenetic profiles for the response of patients to drugs is in progress.
-
Because of the scant amount of published data that describe high-density SNP mapping for the identification of complex susceptibility-gene variants, as well as responses to medicines, examples of SNP-mapping strategies for testing the hypothesis that whole-genome SNP profiling is practical are drawn from work carried out at GlaxoSmithKline.
-
Two examples of adverse-event pharmacogenetics, which highlight the potential speed and utility of these methods in drug development and surveillance, are described.
-
Use of SNP profiling of patients for efficacy pharmacogenetics — with the goal of decreasing the size, complexity, time and expense of clinical trials — is described and contrasted with post-marketing surveillance for adverse-event pharmacogenetic applications.
-
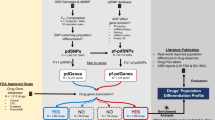
Proposed methodologies for applying SNP-mapping pharmacogenetics for prediction of efficacy and adverse events for individual subjects is summarized.
-
Detailed statistical methods for analysis will be derived empirically, using actual high-density SNP-association data in conjunction with other proposals for statistical evaluation in the literature. Determining a defined pharmacogenetic profile, which contains many variables for high statistical matching, and comparing it with each individual is analogous to current statistical 'forensic' applications of polymorphism screening, in which DNA from two sources can be determined with extraordinary statistical significance.
Abstract
Pharmacogenetic capabilities have changed markedly since The SNP Consortium made a dense single-nucleotide polymorphism (SNP) map freely available in 2001. For more than 40 years, pharmacokinetics and pharmacodynamics of drug-metabolizing molecules were the focus of practical applications. Today, it is possible to use SNP-mapping technologies to create a genetic profile of each individual that can be used to identify patterns of susceptibility genes for common diseases as well as genetic risk/efficacy factors that are related to the effects of drugs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Motulsky, A. J. Pharmacogenetics and ecogenetics in 1991. Pharmacogenetics 1, 2–3 (1991).
Pfsot, D. R., Boyce-Jacino, M. T., & Grant, D. M. A SNPshot: pharmacogenetics and the future of drug therapy. Trends Biotechnol. 18, 334–338 (2000).
Edwards, R. & Aronson, J. K. Adverse drug reactions: definitions, diagnosis, and management. Lancet 356, 1255–1259 (2000).
McLeod, H. L. & Evans, W. E. Pharmacogenomics: unlocking the human genome for better drug therapy. Annu. Rev. Pharmacol. Toxicol. 41, 101–121 (2001).
Sadee, W. Pharmacogenomics: using genetic information to optimize drug therapy. Medscape Pharmacotherapy [online] (cited 10 Jun. 02) (2000).
Kurth, J. H. Pharmacogenomics: future promise of a tool for identifying patients at risk. Drug Info. J. 34, 223–227 (2000).
Roses, A. D. Pharmacogenetics and the practice of medicine. Nature 405, 857–865 (2000).An early review of high-density SNP-mapping technologies, with discussion of ethical considerations of pharmacogenetic SNP mapping compared with disease-gene identification.
Roses, A. D. Pharmacogenetics. Hum. Mol. Genet. 10, 2261–2267 (2001).
Evans, W. E. & Johnson, J. A. Pharmacogenomics: the inherited basis for interindividual differences in drug response. Annu. Rev. Genomics Hum. Genet. 2, 9–39 (2001).
Pirmohamed, M. & Park, B. K. Genetic susceptibility to adverse drug reactions. Trends Pharmacol. Sci. 22, 298–305 (2001).
Fiijal, B. A., Hall, J. M. & Witte, J. S. Clinical trials in the genomic era: effects of protective genotypes on sample size and duration of trial. Control. Clin. Trials 21, 7–20 (2000).
Lazarou, J., Pomeranz, B. H., & Corey, P. N. Incidence of adverse drug reactions in hospitalized patients: a meta-analysis of prospective studies. JAMA 279, 1200–1205 (1998).
Jeffries, D. B., Leakey, D., Lewis, J. A., Payne, S. & Rawlins, M. D. New active substances authorized in the United Kingdom between 1972 and 1994. Br. J. Clin. Pharmacol. 45, 151–156 (1998).
Pirmohamed, M., Breckenridge, A. M., Kitteringham, N. R., & Park, B. K. Adverse drug reactions. BMJ 316, 1295–1298 (1998).
Wieczorek, S. J. & Tsongalis, G. J. Pharmacogenomics: will it change the field of medicine? Clin. Chim. Acta 308, 1–8 (2001).
Lesko, L. J. & Woodcock, J. Pharmacogenomic-guided drug development: regulatory perspective. Pharmacogenomics J. 2, 20–24 (2002).An influential article, which was written by FDA Directors, which invited proposals using SNP-mapping pharmacogenetic technologies, and changed the perception of a widely held view outside the FDA that regulators would not consider such applications.
The SNP Consortium. 〈http://snp.cshl.org/index.html〉 (2002).
Roses, A. D. Pharmacogenetics place in modern medical science and practice. Life Sci. 70, 1471–1480 (2002).
Lai, E., Riley, J., Purvis, I. & Roses, A. D. A 4-Mb high-density single nucleotide polymorphism-based map around human APOE. Genomics 54, 31–38 (1998)The first high-density SNP map that was constructed as a proof of principle for LD mapping around a known susceptibility gene — to test whether the APOE4 polymorphism, which is associated with an increased risk of AD, would have been identified more rapidly by using SNP LD-association mapping.
Roses, A. D. Apolipoprotein E alleles as risk factors in Alzheimer's disease. Annu. Rev. Med. 47, 387–400 (1996).
Farrer, L. A. et al. Effects of age, sex, and ethnicity on the association between apolipoprotein E genotype and Alzheimer disease. A meta-analysis. APOE and Alzheimer Disease Meta Analysis Consortium. JAMA 278, 1349–1356 (1997).
Tang, M. X. et al. The APOE ɛ4 allele and the risk of Alzheimer disease among African Americans, whites, and Hispanics. JAMA 279, 751–755 (1998).
Graff-Radford, N. R. et al. Association between apolipoprotein E genotype and Alzheimer disease in African American subjects. Arch. Neurol. 59, 594–600 (2002).
Kardaun, J. W. et al. Genotypes and phenotypes for apolipoprotein E and Alzheimer disease in the Honolulu–Asia aging study. Clin. Chem. 46, 1548–1554 (2000).
Martin, E. R. et al. SNPing away at complex diseases: analysis of single-nucleotide polymorphisms around APOE in Alzheimer disease. Am. J. Hum. Genet. 67, 383–394 (2000).A detailed analysis of the first high-density SNP map around the APOE gene, which was carried out as a proof of principle in reference 19.
Martin, E. R. et al. Analysis of association at SNPs in the APOE region. Genomics 63, 7–12 (2000).
McCarthy, L. C. et al. Single-nucleotide polymorphism alleles in the insulin receptor gene are associated with typical migraine. Genomics 78, 135–149 (2001).The insulin receptor was the first susceptibility gene for a complex disease that was identified using high-density SNP mapping of a genetic-linkage region.
Hewett, D. et al. Identification of a psoriasis susceptibility candidate gene by linkage disequilibrium mapping with a localized single nucleotide polymorphism map. Genomics 79, 305–314 (2002).This research identifies a gene that contains specific SNP associations with psoriasis, and defines a gene that was not a previous disease susceptibility candidate.
Rioux, J. D. et al. Genetic variation in the 5q31 cytokine gene cluster confers susceptibility to Crohn disease. Nature Genet. 29, 223–228 (2001).The identification of a region using SNP mapping that contains a gene cluster that is associated with susceptibility to Crohn's disease.
Bergstresser, P. R. Sensitization and elicitation of inflammation in contact dermatitis. Immunol. Ser. 46, 219–245 (1989).
Fleming, C. J., Burden, A. D. & Forsyth, A. The genetics of allergic contact hypersensitivity to nickel. Contact Dermatitis 41, 251–253 (1999).
Ardlie, K. G., Kruglyak, L. & Seielstad, M. Patterns of linkage disequilibrium in the human genome. Nature Rev. Genet. 3, 299–309 (2002).
Cantor, C. Whole genome SNP scans: the reality. The Human Genome Organisation [online] (cited 31 May 2002) (2002).
Foster, R. H. & Foulds, D. Abacavir. Drugs 55, 729–736 (1998).
Symonds, W. et al. Risk factor analysis for hypersensitivity reactions to abacavir. Clin. Ther. (in the press).
Hetherington, S. W. et al. Hypersensitivity reactions during therapy with abacavir: analysis of 636 cases for clinical presentation and risk factors. [online] (cited 31 May 2002) 7th Conference on Retroviruses and Opportunistic Infections (2000).
Brew, B. J. et al. Phase III, randomized, double-blind, placebo controlled, multicentre study to evaluate the safety and efficacy of abacavir (ABC 1592) in HIV-1 infected subjects with AIDS dementia complex (CNA3001). 12th World AIDS Conference Abstract 32,192 (1998).
Brinkman, K., Smeitink, J. A., Romijn, A., & Reiss, P. Mitochondrial toxicity induced by nucleoside-analogue reverse transcriptase inhibitors is a key factor in the pathogenesis of antiretroviral-therapy-related lipodystrophy. Lancet 354, 1112–1115 (1999).
Hetherington, S. et al. Genetic variations in HLA-B region and hypersensitivity reactions to abacavir. Lancet 359, 1121–1122 (2002).This paper shows that several polymorphisms in a region of extended LD confer susceptibility to the HSR to abacavir, using a list of candidate genes that are thought to be involved in immunological disorders. By chance, SNPs from two distinct genes were in extended LD, and so defined a larger-than-expected region, which shows that estimates of the number of SNPs that are needed for whole-genome screening profiles might be too large.
Mallal, S. et al. Association between the presence of HLA-B*5701, HLA-DR7, and HLA-GQ3 and hypersensitivity to HIV-1 reverse-transcriptase inhibitor abacavir. Lancet 359, 727–732 (2002).These independent experiments look at specific HLA regions as candidate sets of polymorphisms for association to the HSR to abacavir and, along with reference 39 , provide immediate published confirmation of this LD region.
Monaghan, G., Ryan, M., Seddon, H. R. & Burchell, B. Genetic variation in bilirubin UPD-glucuronosyltransferase gene promoter and Gilbert's syndrome. Lancet 347, 578–581 (1996).
Aono, S. et al. Analysis of genes for bilirubin UPD-glucuronosyltransferase in Gilbert's syndrome. Lancet 345, 958–959 (1995).
Bosma, P. J. et al. The genetic basis of the reduced expression of bilirubin UDP-glucuronosyltransferase 1 in Gilbert's syndrome. N. Engl. J. Med. 333, 1171–1175 (1995).
Bosma, P. J. et al. Sequence of exons and the flanking region of human bilirubin–UDP-glucuronosyltransferase gene complex and identification of a genetic mutation in a patient with Crigler–Najjar syndrome, type I. Hepatology 15, 941–947 (1992).
Roses, A. D. Pharmacogenetics and future drug development and delivery. Lancet 355, 1358–1361 (2000).
Petricoin, E. F. et al. Use of proteomic patterns in serum to identify ovarian cancer. Lancet 359, 572–577 (2002).
Buchanan, A. et al. Pharmacogenetics: ethical issues and policy options. Kennedy Inst. Ethics J. (in the press).
Robertson, J. A. Pharmacogenetic challenges for the health care system. Health Affairs (in the press).
Wolf, C. R., Smith, G. & Smith, R. L. Science, medicine and the future: pharmacogenetics. BMJ 320, 987–990 (2000).
Ingelman-Sundberg, M. Pharmacogenetics: an opportunity for a safer and more efficient pharmacotherapy. J. Int. Med. 250, 186–200 (2001).
Park, K. B. & Pirmohamed, M. Toxicogenetics in drug development. Toxicol. Lett. 120, 281–291 (2001).
Cardon, L. R. et al. Testing drug response in the presence of genetic information: sampling issues for clinical trials. Pharmacogenetics 10, 503–510 (2000).
Acknowledgements
A.D.R. acknowledges the GlaxoSmithKline (GSK) department heads and project leaders whose teams contributed to the experiments and examples described in this review: D. Burns, D. Campbell, R. Dement, A. Hughes, E. Lai, P. Manasco, A. McCarthy, I. Purvis, J. Riley, A. Saunders, S. Sehgal, Y. Smithies and N. Spurr. Also, the many scientists involved in creating the SNP mapping strategy and capability within GSK; T. Isenhour and E. McPherson, who assisted in preparing and editing the manuscript; A. Holden and D. Wang, who accelerated The SNP Consortium to be on time, under budget and to exceed all expectations; and M. McPeak, E. Haner, M. Beadle and S. Groh, who contributed support for team coordination. J. Niedel, T. Yamada, A. Hennah, J. Palmer, J. Robinson, B. Koch and K. Spitz provided essential early and continued support for pharmacogenetics and pharmacogenomics in the Genetics Research Directorate in GSK.
Author information
Authors and Affiliations
Related links
Related links
DATABASES
Cancer.gov
LocusLink
Medscape DrugInfo
OMIM
Crigler–Najjar syndrome type I
FURTHER INFORMATION
LINKS
Glossary
- SINGLE-NUCLEOTIDE POLYMORPHISM
-
(SNP). A specific location in a DNA sequence at which different people have different DNA bases. This can change the protein sequence, leading to disease (for example, sickle-cell disease), or have no known consequences.
- SNP MAPPING
-
A linear map of SNPs across the genome allows genetic traits to be localized by statistical association to the specific region of the genome that is marked by the SNP or several SNPs nearby. In the cases of complex diseases or responses to medicine, multiple SNP markers in several specific regions of the genome act as markers that are associated with the phenotype (disease or drug response).
- CANDIDATE-GENE ASSOCIATIONS
-
A candidate gene is frequently indicated by previous hypotheses or genetic-linkage studies. Gene variants can cause or enhance susceptibility to a disease or drug-response phenotype.
- SNP-LINKAGE MAPPING PANEL
-
The collection of SNPs that defines the linear map of the genome that is being studied. To localize the APOE gene, these SNPs would be located on chromosome 19. For HLA-B57, the SNPs are located on chromosome 6. To find other unknown regions, a SNP mapping panel that covers the whole genome can be used.
- LINKAGE DISEQUILIBRIUM
-
(LD). Defines a region of allele (or inherited-variation) sharing along a relatively small length of the genome. If there is an association with one marker, then the likelihood of association with another marker within that region is increased.
- PENETRANT GENETIC DISEASES
-
Describes the expression of the phenotype that results from the inheritance of disease-associated or phenotype-associated alleles. Some diseases have relatively highly penetrant mutant alleles, such as Huntington's disease. Penetrance of other traits, such as eye colour or late-onset Alzheimer's disease, might be due to combinations of less-penetrant polymorphic alleles found at several locations along the genome.
- AUTOSOMAL RECESSIVE
-
A mode of inheritance that requires that the mutation of a single gene is present on both paternally and maternally derived alleles for the clinical phenotype to be expressed.
- AUTOSOMAL DOMINANT
-
A mode of inheritance that requires that the mutation of a single gene is present on either of the paternally and maternally derived alleles for the clinical phenotype to be expressed.
- SNP PRINTsm
-
Describes the pattern of inheriting SNP variations from a panel of SNPs. In the case of SNP mapping along the whole genome, it is analogous to the detection of highly private sets of traits, such as those that determine fingerprints, which make individuals unique. In the case of pharmacogenetics, it is the specific set of measured SNPs that define a drug response, and which can therefore be abstracted from the whole map to develop a more simple medicine-response profile.
- MEDICINE-RESPONSE PROFILE
-
(MRP). A test or set of tests that indicates the likely response of a patient to a medicine. In the context of developing a test from a SNP Printsm, the test might be the selection of SNPs from the genome scan that are associated with the phenotypic response; for example, adverse event.
- HYPERBILIRUBINAEMIA
-
Bilirubin is a metabolite of cholesterol, and its elevation above normal serum levels is called hyperbilirubinaemia. This state can be caused by several liver- and gall-bladder-related problems.
- CLUSTER ANALYSIS
-
A formal method of statistical analysis that can be used to determine the significance of clusters of data points. For example, the clustering of disease-associated SNP variants in a region of LD, or the identification of multiple regions of nearby SNP variants (∼LD) at particular locations along the genome that defines a high probability that a specific drug response will occur.
Rights and permissions
About this article
Cite this article
Roses, A. Genome-based pharmacogenetics and the pharmaceutical industry. Nat Rev Drug Discov 1, 541–549 (2002). https://doi.org/10.1038/nrd840
Issue Date:
DOI: https://doi.org/10.1038/nrd840
This article is cited by
-
Generating Genome-Scale Candidate Gene Lists for Pharmacogenomics
Clinical Pharmacology & Therapeutics (2009)
-
Risk in CNS drug discovery: focus on treatment of Alzheimer's Disease
BMC Neuroscience (2008)
-
Pharmacogenetics in drug discovery and development: a translational perspective
Nature Reviews Drug Discovery (2008)
-
How can we realize the promise of personalized antidepressant medicines?
Nature Reviews Neuroscience (2008)
-
Complex disease-associated pharmacogenetics: drug efficacy, drug safety, and confirmation of a pathogenetic hypothesis (Alzheimer's disease)
The Pharmacogenomics Journal (2007)